Functions converting between phylogenetic trees and their unique decimal representation, based on a concept by John Tromp, employed in Li et al. (1996) .
Usage
as.TreeNumber(x, ...)
# S3 method for class 'phylo'
as.TreeNumber(x, ...)
# S3 method for class 'multiPhylo'
as.TreeNumber(x, ...)
# S3 method for class 'character'
as.TreeNumber(x, nTip, tipLabels = TipLabels(nTip), ...)
# S3 method for class 'TreeNumber'
as.TreeNumber(x, ...)
# S3 method for class 'MixedBase'
as.TreeNumber(x, ...)
# S3 method for class 'TreeNumber'
as.MixedBase(x, ...)
# S3 method for class 'integer64'
as.MixedBase(x, tipLabels = NULL, ...)
# S3 method for class 'numeric'
as.MixedBase(x, tipLabels = NULL, ...)
# S3 method for class 'numeric'
as.phylo(x, nTip = attr(x, "nTip"), tipLabels = attr(x, "tip.label"), ...)
# S3 method for class 'TreeNumber'
as.phylo(x, nTip = attr(x, "nTip"), tipLabels = attr(x, "tip.label"), ...)
as.MixedBase(x, ...)
# S3 method for class 'MixedBase'
as.MixedBase(x, ...)
# S3 method for class 'phylo'
as.MixedBase(x, ...)
# S3 method for class 'multiPhylo'
as.MixedBase(x, ...)
# S3 method for class 'MixedBase'
as.phylo(x, nTip = attr(x, "nTip"), tipLabels = attr(x, "tip.label"), ...)Arguments
- x
Integer identifying the tree (see details).
- ...
Additional parameters for consistency with S3 methods (unused).
- nTip
Integer specifying number of leaves in the tree.
- tipLabels
Character vector listing the labels assigned to each tip in a tree, perhaps obtained using
TipLabels().
Value
as.TreeNumber() returns an object of class TreeNumber with
attributes nTip and tip.label.
For trees with at most 19 leaves the underlying storage is a single
integer64 value (class c("TreeNumber", "integer64")), enabling
integer64 arithmetic and exact round-tripping through as.MixedBase().
For trees with 20–51 leaves the number exceeds 2^64, so it is stored as a
decimal character string (class c("TreeNumber", "character")).
If x is a list of trees or a multiPhylo object,
as.TreeNumber() returns a corresponding list of TreeNumber objects.
as.phylo.numeric() returns a tree of class phylo.
Details
There are NUnrooted(n) unrooted trees with n leaves.
As such, each n-leaf tree can be uniquely identified by a non-negative
integer x < NUnrooted(n).
This integer can be converted by a tree by treating it as a mixed-base number, with bases 1, 3, 5, 7, … (2 n - 5).
Each digit of this mixed base number corresponds to a leaf, and determines the location on a growing tree to which that leaf should be added.
We start with a two-leaf tree, and treat 0 as the origin of the tree.
We add leaf 2 by breaking an edge and inserting a node (numbered
2 + nTip - 1).
In this example, we'll work up to a six-leaf tree; this node will be numbered
2 + 6 - 1 = 7.
There is only one edge on which leaf 2 can be added. Let's add node 7 and
leaf 2:
There are now three edges on which leaf 3 can be added. Our options are:
Option 0: the edge leading to 1;
Option 1: the edge leading to 2;
Option 2: the edge leading to 7.
If we select option 1, we produce:
1 is now the final digit of our mixed-base number.
There are five places to add leaf 4:
Option 0: the edge leading to 1;
Option 1: the edge leading to 2;
Option 2: the edge leading to 3;
Option 3: the edge leading to 7;
Option 4: the edge leading to 8.
If we chose option 3, then 3 would be the penultimate digit of our
mixed-base number.
If we chose option 0 for the next two additions, we could specify this tree with the mixed-base number 0021. We can convert this into decimal:
0 × (1 × 3 × 5 × 9) +
0 × (1 × 3 × 5) +
3 × (1 × 3) +
1 × (1)
= 10
as.TreeNumber() supports up to 51 leaves.
For trees with at most 19 leaves, the number fits in a 64-bit integer and
the TreeNumber is stored as an integer64 (via the bit64 package),
enabling arithmetic and exact round-tripping via as.MixedBase().
For trees with 20–51 leaves, there are more than 2^64 distinct topologies,
so the tree number is stored as a decimal character string instead.
Package developers can use the C++ header TreeTools/tree_number.h
(via LinkingTo: TreeTools) for the underlying 256-bit encoding
(tree_num_t) directly.
References
Li M, Tromp J, Zhang L (1996). “Some notes on the nearest neighbour interchange distance.” In Goos G, Hartmanis J, Leeuwen J, Cai J, Wong CK (eds.), Computing and Combinatorics, volume 1090, 343–351. Springer, Berlin, Heidelberg. ISBN 978-3-540-61332-9. doi:10.1007/3-540-61332-3_168 .
See also
Describe the shape of a tree topology, independent of leaf labels:
TreeShape()
Other tree generation functions:
ConstrainedNJ(),
GenerateTree,
NJTree(),
TrivialTree
Other 'TreeNumber' utilities:
is.TreeNumber(),
print.TreeNumber()
Examples
tree <- as.phylo(10, nTip = 6)
plot(tree)
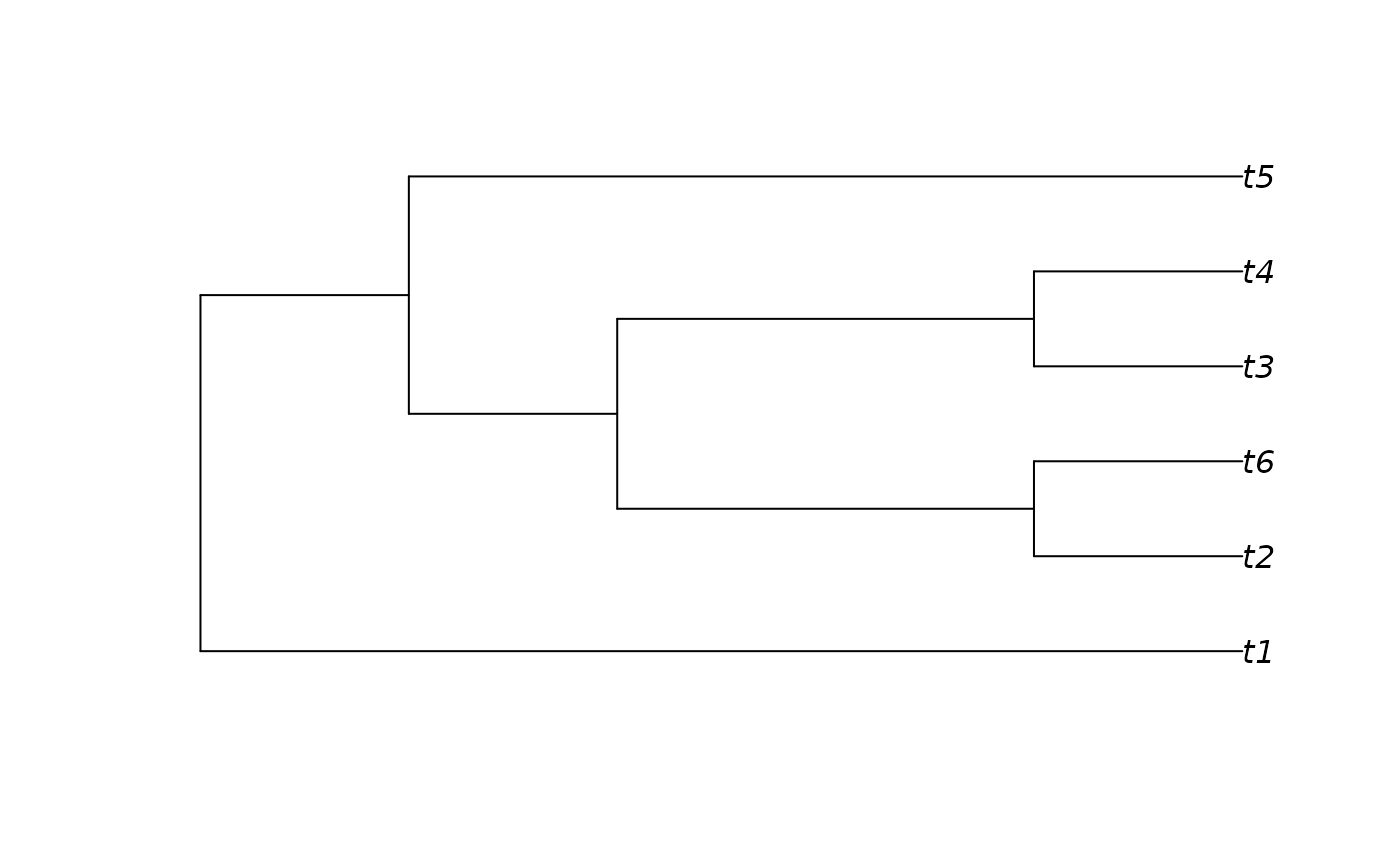 as.TreeNumber(tree)
#> Phylogenetic tree number 10 of 105
#> 6 tips: t1 t2 t3 t4 t5 t6
# Trees with 20--51 leaves are stored as decimal strings:
as.TreeNumber(BalancedTree(19)) # integer64-backed
#> Phylogenetic tree number 3259279213732796827 of 6332659870762850625
#> 19 tips: t1 t2 t3 t4 t5 t6 t7 t8 t9 t10 t11 t12 t13 t14 t15 t16 t17 t18 t19
as.TreeNumber(BalancedTree(51)) # character-backed
#> Phylogenetic tree number 27388803913187622481001283703331864424119472007335608662299812499724470715408 of 2.752921e+76
#> 51 tips: t1 t2 t3 t4 t5 t6 t7 t8 t9 t10 t11 t12 t13 t14 t15 t16 t17 t18 t19 t20 t21 t22 t23 t24 t25 t26 t27 t28 t29 t30 t31 t32 t33 t34 t35 t36 t37 t38 t39 t40 t41 t42 t43 t44 t45 t46 t47 t48 t49 t50 t51
# If > 9 digits, represent the tree number as a string.
treeNumber <- as.TreeNumber("1234567890123", nTip = 14)
tree <- as.phylo(treeNumber)
as.phylo(0:2, nTip = 6, tipLabels = letters[1:6])
#> 3 phylogenetic trees
as.TreeNumber(tree)
#> Phylogenetic tree number 10 of 105
#> 6 tips: t1 t2 t3 t4 t5 t6
# Trees with 20--51 leaves are stored as decimal strings:
as.TreeNumber(BalancedTree(19)) # integer64-backed
#> Phylogenetic tree number 3259279213732796827 of 6332659870762850625
#> 19 tips: t1 t2 t3 t4 t5 t6 t7 t8 t9 t10 t11 t12 t13 t14 t15 t16 t17 t18 t19
as.TreeNumber(BalancedTree(51)) # character-backed
#> Phylogenetic tree number 27388803913187622481001283703331864424119472007335608662299812499724470715408 of 2.752921e+76
#> 51 tips: t1 t2 t3 t4 t5 t6 t7 t8 t9 t10 t11 t12 t13 t14 t15 t16 t17 t18 t19 t20 t21 t22 t23 t24 t25 t26 t27 t28 t29 t30 t31 t32 t33 t34 t35 t36 t37 t38 t39 t40 t41 t42 t43 t44 t45 t46 t47 t48 t49 t50 t51
# If > 9 digits, represent the tree number as a string.
treeNumber <- as.TreeNumber("1234567890123", nTip = 14)
tree <- as.phylo(treeNumber)
as.phylo(0:2, nTip = 6, tipLabels = letters[1:6])
#> 3 phylogenetic trees